Research Project:Development of Antibody-Like Proteins Using TRAP Display
What is antibody-like protein?
One of the therapeutic approaches for diseases is antibody-based drugs. Antibodies are a type of protein called immunoglobulins that recognize and bind to specific targets. For example, antibody therapeutics targeting proteins on the surface of cancer cells can bind to these target proteins and induce the death of cancer cells. Antibody drugs are characterized by high therapeutic efficacy and relatively low side effects, and they are expected to be effective treatments for intractable diseases for which no established therapies exist. However, antibodies are large and structurally complex, making them costly to produce. Therefore, in recent years, “small-sized antibodies” have attracted attention as alternative tools to conventional antibodies. These small-sized antibodies are proteins smaller than immunoglobulins that can bind to specific targets like antibodies. So we refer to them as antibody-like proteins.
What is TRAP display?
To date, many types of antibody-like proteins have been reported. Representative examples include Monobodies [1], which are based on fibronectin, and Anticalins [2], which are based on lipocalins. To create antibody-like proteins, it is necessary to select a protein scaffold and engineer it to bind to a target of interest. Proteins are composed of chains of amino acids, and they can be modified by replacing certain amino acids with others. However, since there are 20 canonical amino acids, it is difficult to determine which amino acid substitutions—and at which positions—will result in desired binding properties. Therefore, an effective strategy is to generate a large number of antibody-like protein candidates by introducing random mutations at various positions and then select those that bind to the target (screening or selection).
mRNA display, first reported in 1997 [3, 4], is a technique that enables the selection of desired antibody-like proteins from libraries containing up to 10^13 variants. However, this method has drawbacks, including procedural complexity and long processing times. To address these issues, we developed TRanscription-TRanslation coupled with Association of Puromycin-linker (TRAP) display, a simplified and accelerated version of mRNA display. Currently, we are using TRAP display to generate antibody-like proteins against a variety of targets.
Overview of Important Papers
T. Ishizawa, T. Kawakami, P. C. Reid, H. Murakami*, TRAP display: a high-speed selection method for the generation of functional polypeptides.
J. Am. Chem. Soc. 2013, 135, 5433-5440.
https://doi.org/10.1021/ja312579u
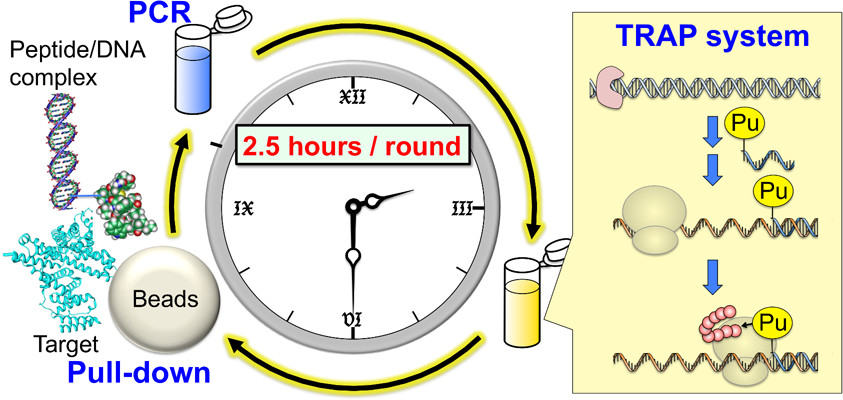
TRAP (TRanscription–TRanslation coupled with Association of Puromycin-linker) display was developed as a simplified and accelerated version of mRNA display. This study reported the selection of cyclic peptides that bind to specific targets.
While peptide selection by mRNA display typically requires 2–3 weeks, TRAP display enables the process to be completed within 1–2 days. The development of TRAP display is expected to accelerate peptide selection and contribute to the discovery of various functional molecules.
T. Kondo, Y. Iwatani, K. Matsuoka, T. Fujino, S. Umemoto, Y. Yokomaku, K. Ishizaki, S. Kito, T. Sezaki, G. Hayashi, H. Murakami*, Antibody-like proteins that capture and neutralize SARS-CoV-2.
Sci. Adv., 2020, 6(42), eabd3916.
https://doi.org/10.1126/sciadv.abd3916
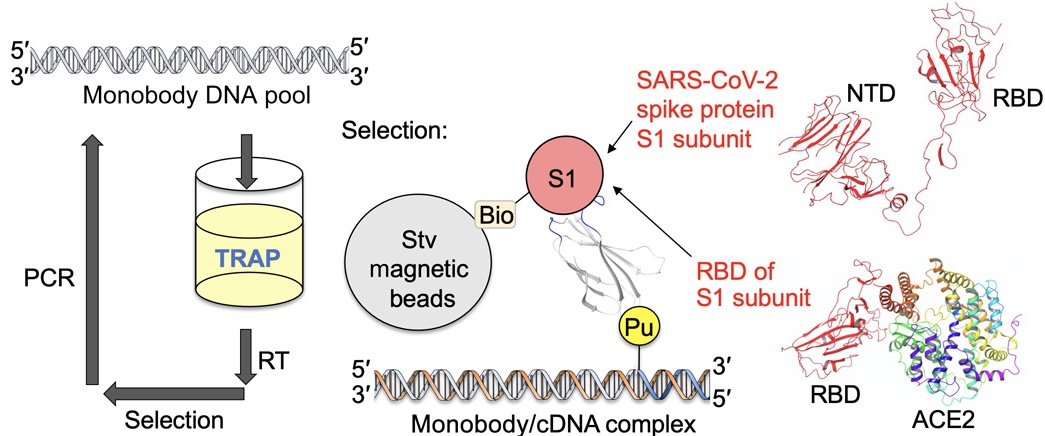
TRAP display was further improved to enable the selection of antibody-like proteins. Using this method, Monobodies targeting SARS-CoV-2 were successfully generated.
We focused on the receptor-binding domain (RBD) of SARS-CoV-2. The RBD is part of the spike protein on the viral surface and is known to bind to angiotensin-converting enzyme 2 (ACE2) on human cells. Therefore, monobodies that bind to the RBD can block viral entry into host cells and inhibit infection.
As a result of TRAP display selection, high-affinity monobodies (dissociation constant: 0.42–1.9 nM) were obtained within only 4 days. TRAP display is a powerful technology that enables rapid generation of artificial antibodies against various targets, including emerging viruses.
S. Umemoto, T. Kondo, T. Fujino, G. Hayashi, H. Murakami*, Large-scale analysis of mRNA sequences localized near the start and amber codons and their impact on the diversity of mRNA display libraries.
Nucleic Acids Res., 2023, 51, 7465-7479.
https://doi.org/10.1093/nar/gkad555
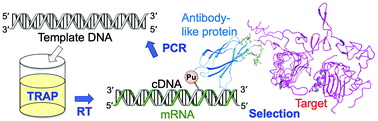
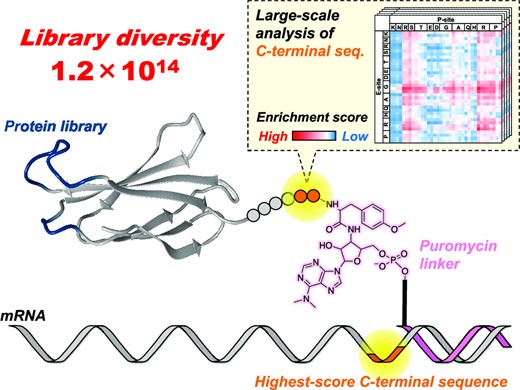
In TRAP display, a DNA library is first constructed by introducing random mutations into sequences encoding antibody-like proteins. When this library is added to the TRAP system, transcription from DNA to mRNA, conjugation with puromycin, and translation into antibody-like proteins proceed continuously. Because puromycin forms a covalent bond with the C-terminus of the translated polypeptide, mRNA/puromycin–polypeptide complexes are generated. This enables a direct linkage between antibody-like proteins and their encoding mRNAs, allowing sequence identification of binders.
Therefore, maintaining this correspondence is crucial. Antibody-like proteins not linked to mRNA cannot provide sequence information even if recovered. Increasing the number of properly linked complexes expands the searchable library diversity and improves the binding affinity of obtaining binders [5]. In this study, improving the formation efficiency of mRNA/puromycin–polypeptide complexes enhanced the overall selection efficiency of TRAP display.
T. Kondo, M. Eguchi, S. Kito, T. Fujino, G. Hayashi, H. Murakami.*, cDNA TRAP display for rapid and stable in vitro selection of antibody-like proteins.
Chem. Commun., 2021, 57, 2416-2419.
https://doi.org/10.1039/D0CC07541HAs described above, maintaining the linkage between antibody-like proteins and mRNA is essential in TRAP display. However, mRNA is susceptible to degradation by RNases, which are ubiquitous in the environment, including on human skin, hair, and in biological samples such as cells and serum. This makes contamination likely and limits applications under RNase-rich conditions.
cDNA display was developed to overcome this issue by enabling selection under RNase-containing conditions. Following this strategy, we developed cDNA TRAP display, which allows rapid and stable selection under a wider range of conditions.
G. Hayashi*, T. Naito, S. Miura, N. Iwamoto, Y. Usui, M. Bando-Shimizu, S. Suzuki, K. Higashi, M. Nonaka*, S. Oishi*, H. Murakami*, Generating a mirror-image monobody targeting MCP-1 via TRAP display and chemical protein synthesis.
Nat. Commun., 2024, 15, 10723.
https://doi.org/10.1038/s41467-024-54902-xIn TRAP display, antibody-like proteins are typically synthesized using L-amino acids through translation. However, since endogenous proteins are also composed of L-amino acids, such proteins are prone to degradation and immune recognition (immunogenicity). In contrast, D-amino acid-based (mirror-image) proteins are less susceptible to these issues.
However, direct synthesis of D-proteins via translation is difficult. To address this, a mirror-image strategy has been proposed: a target protein is chemically synthesized in its D-form, L-form proteins that bind to this D-target are selected, and then these proteins are chemically synthesized in the D-form. This results in D-proteins that bind to the natural L-target [9].
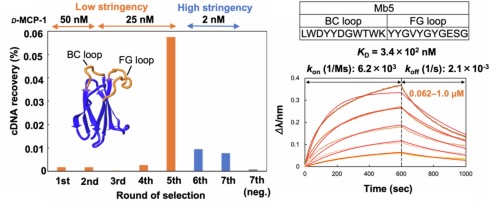
We applied this strategy to TRAP display. After selecting an L-form monobody against D-MCP-1, we chemically synthesized its D-form, which binds to natural L-MCP-1. This approach is expected to accelerate the development of highly effective therapeutics based on D-proteins.
References
[1] A. Koide, C. W. Bailey, X. Huang, S. Koide, The fibronectin type III domain as a scaffold for
novel binding proteins. J. Mol. Biol., 1998, 284, 1141-1151.
[2] G. Beste, F. S. Schmidt, T. Stibora, A. Skerra, Small antibody-like proteins with prescribed
ligand specificities derived from the lipocalin fold. Proc. Natl. Acad. Sci. USA, 1999, 96, 1898-1903.
[3] R. W. Roberts, J. W. Szostak, RNA-peptide fusions for the in vitro selection of peptides and
proteins. Proc. Natl. Acad. Sci. USA, 1997, 94, 12297-12302.
[4] N. Nemoto, E. Miyamoto-Sato, Y. Husimi, H. Yanagawa, In vitro virus: Bonding of mRNA bearing
puromycin at the 3'-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro.
FEBS Lett. 1997, 414, 405-408.
[5] G. Kamalinia, B. J. Grindel, T. T. Takahashi, S. W. Millward, R. W. Roberts, Directing evolution
of novel ligands by mRNA display. Chem. Soc. Rev. 2021, 50, 9055-9103.
[6] M. Kurz, K. Gu, A. Al-Gawari, P. A. Lohse, cDNA-protein fusions: Covalent protein–gene conjugates for the in vitro selection of peptides and proteins. ChemBioChem, 2001, 2, 666–672.
[7] I. Tabuchi, S. Soramoto, N. Nemoto, Y. Husimi, An in vitro DNA virus for in vitro protein evolution. FEBS Lett. 2001, 508, 309–312.
[8] J. Yamaguchi, M. Naimuddin, M. Biyani, T. Sasaki, M. Machida, T. Kubo, T. Funatsu, Y. Husimi, N. Nemoto, cDNA display: a novel screening method for functional disulfide-rich peptides by solid-phase synthesis and stabilization of mRNA–protein fusions. Nucleic Acids Res. 2009, 37, e108.
[9] T. N. M. Schumacher, L. M. Mayr, D. L. Minor, Jr., M. A. Milhollen, M. W. Burgess, P. S. Kim, Identification of d-Peptide Ligands Through Mirror-Image Phage Display. Science, 1996, 271, 1854-1857.